Map
Halie
7/13/2021
4. Map
4.1 Read in Spatial Data
4.1.1 Install packages
# require() is like library() except returns FALSE if missing (vs error)
if(!require(librarian)){
install.packages("librarian")
library(librarian)
library(Rcpp)
}## Loading required package: librarian#librarian::shelf() is like library() except installs package if missing, even from Github if include owner/repo
shelf(NOAA-EDAB/ecodata, sf)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.4.2 Get spatial data
epu_sf <- ecodata::epu_sf %>%
st_transform(4326)So we see a geometry list column.
class(epu_sf)## [1] "sf" "data.frame"g1 <- epu_sf$geometry[1]
# see in Environment pane, expand g1
plot(epu_sf)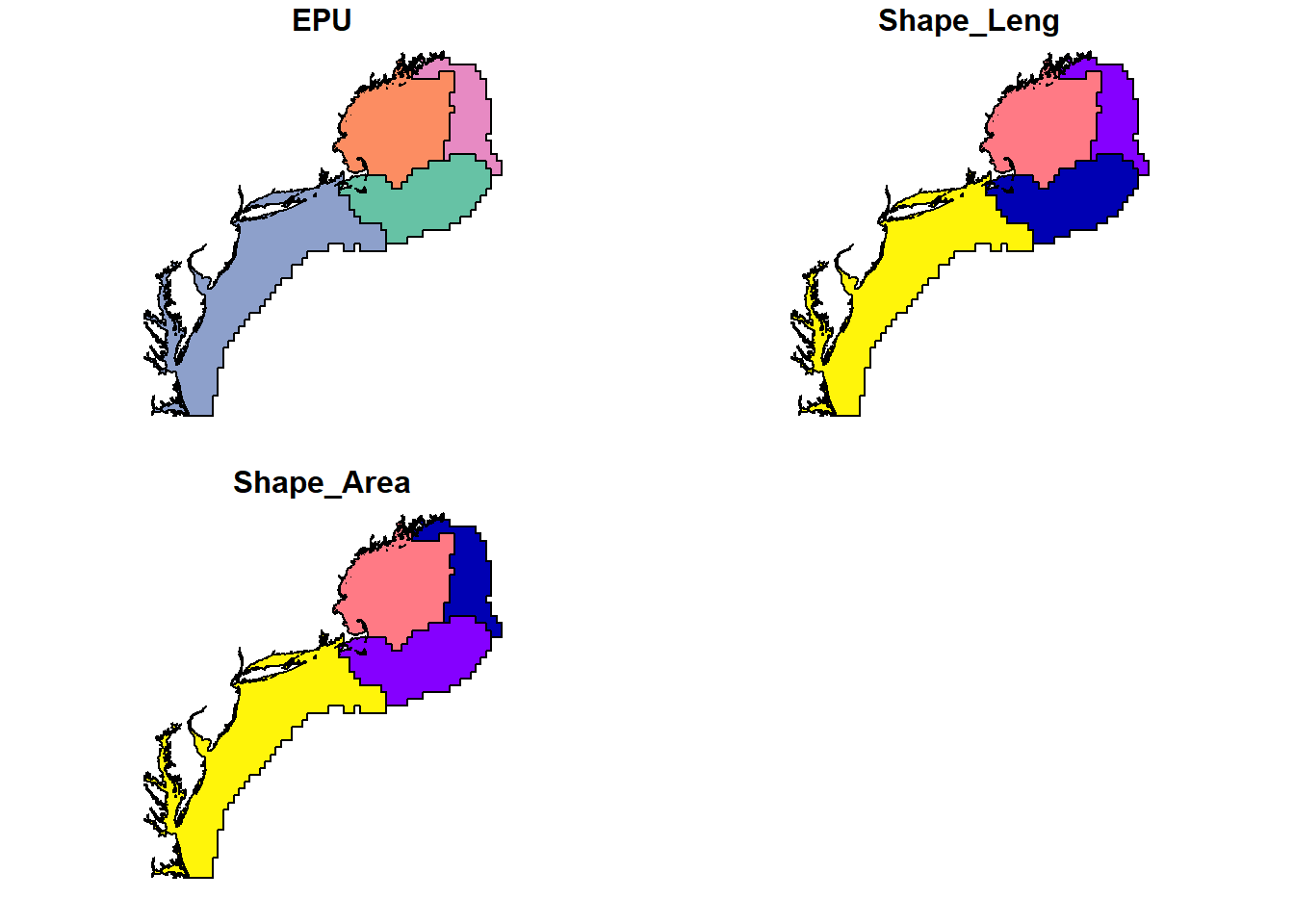
plot(epu_sf["EPU"])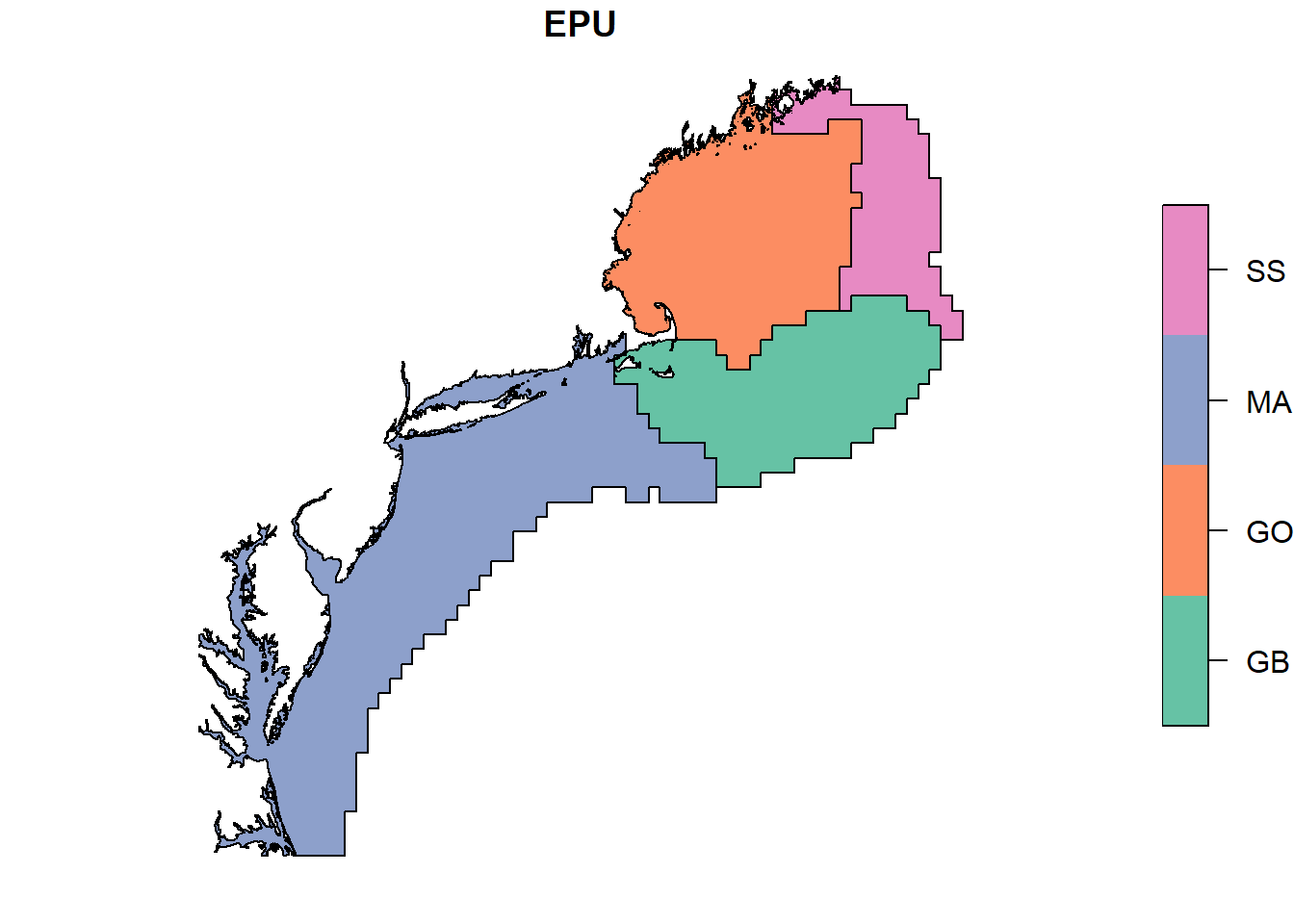
shelf(mapview)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.mapview(epu_sf)shelf(leaflet)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.leaflet() %>%
#addTiles() %>%
addProviderTiles(providers$Esri.OceanBasemap) %>%
addPolygons(data = epu_sf)3.3 Group by
sf is “tidy”
3.4 Extract from ERDDAP
ERDDAP - Multi-scale Ultra-high Resolution (MUR) SST Analysis fv04.1, Global, 0.01 degrees, 2002-present ,Monthly- Data Access form (https://coastwatch.pfeg.noaa.gov/erddap/griddap/jplMURSST41mday.html)
CoastWatch ERDDAP (https://coastwatch.pfeg.noaa.gov/erddap/index.html): search for “SST”:
- jplMURSST41mday: ERDDAP - Multi-scale Ultra-high Resolution (MUR) SST Analysis fv04.1, Global, 0.01°, 2002-present, Monthly - Data Access Form (https://coastwatch.pfeg.noaa.gov/erddap/griddap/jplMURSST41mday.html)
shelf(here, rerddap)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.sst_gd_rds <- here("data/sst_gd.rds")
epu_bb <- st_bbox(epu_sf)
epu_bb## xmin ymin xmax ymax
## -77.00000 35.83270 -65.66667 44.66667sst_info <- info('jplMURSST41mday')
sst_info## <ERDDAP info> jplMURSST41mday
## Base URL: https://upwell.pfeg.noaa.gov/erddap/
## Dataset Type: griddap
## Dimensions (range):
## time: (2002-06-16T00:00:00Z, 2021-06-16T00:00:00Z)
## latitude: (-89.99, 89.99)
## longitude: (-179.99, 180.0)
## Variables:
## mask:
## nobs:
## sst:
## Units: degree_Cif(!file.exists(sst_gd_rds)){
sst_gd <- griddap(sst_info, fields = "sst", time = c("2020-06-16", "2021-06-16"),longitude = epu_bb[c("xmin", "xmax")], latitude = epu_bb[c("ymin", "ymax")])
saveRDS(sst_gd,file = sst_gd_rds)
}
sst_gd <- readRDS(sst_gd_rds)
sst_gd## <ERDDAP griddap> jplMURSST41mday
## Path: [C:\Users\HALIE~1.OFA\AppData\Local\Cache\R\rerddap\6d0dbbf4f950f8dc4ffca5dd9b2cf661.nc]
## Last updated: [2021-07-14 11:35:29]
## File size: [52.2 mb]
## Dimensions (dims/vars): [3 X 1]
## Dim names: time, latitude, longitude
## Variable names: Sea Surface Temperature Monthly Mean
## data.frame (rows/columns): [13046670 X 4]
## # A tibble: 13,046,670 x 4
## time lat lon sst
## <chr> <dbl> <dbl> <dbl>
## 1 2020-06-16T00:00:00Z 35.8 -77 NA
## 2 2020-06-16T00:00:00Z 35.8 -77.0 NA
## 3 2020-06-16T00:00:00Z 35.8 -77.0 NA
## 4 2020-06-16T00:00:00Z 35.8 -77.0 NA
## 5 2020-06-16T00:00:00Z 35.8 -77.0 NA
## 6 2020-06-16T00:00:00Z 35.8 -76.9 NA
## 7 2020-06-16T00:00:00Z 35.8 -76.9 NA
## 8 2020-06-16T00:00:00Z 35.8 -76.9 NA
## 9 2020-06-16T00:00:00Z 35.8 -76.9 NA
## 10 2020-06-16T00:00:00Z 35.8 -76.9 NA
## # ... with 13,046,660 more rowsnames(sst_gd)## [1] "summary" "data"shelf(dplyr, ggplot2, mapdata)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.#coastline
coast <- map_data("worldHires", xlim = epu_bb[c("xmin", "xmax")], ylim = epu_bb[c("ymin", "ymax")], lforce = "e")
sst_df_last <- sst_gd$data %>%
filter(time == max(time))
# summary(sst_last)
ggplot(data = sst_df_last, aes(x = lon, y = lat, fill = sst)) + geom_polygon(data = coast, aes(x = long, y = lat, group = group), fill = "grey80") + geom_tile() + scale_fill_gradientn(colors = rerddap::colors$temperature, na.value = NA) + theme_bw() + ylab("Latitude") + xlab("Longitude") + ggtitle("Latest SST")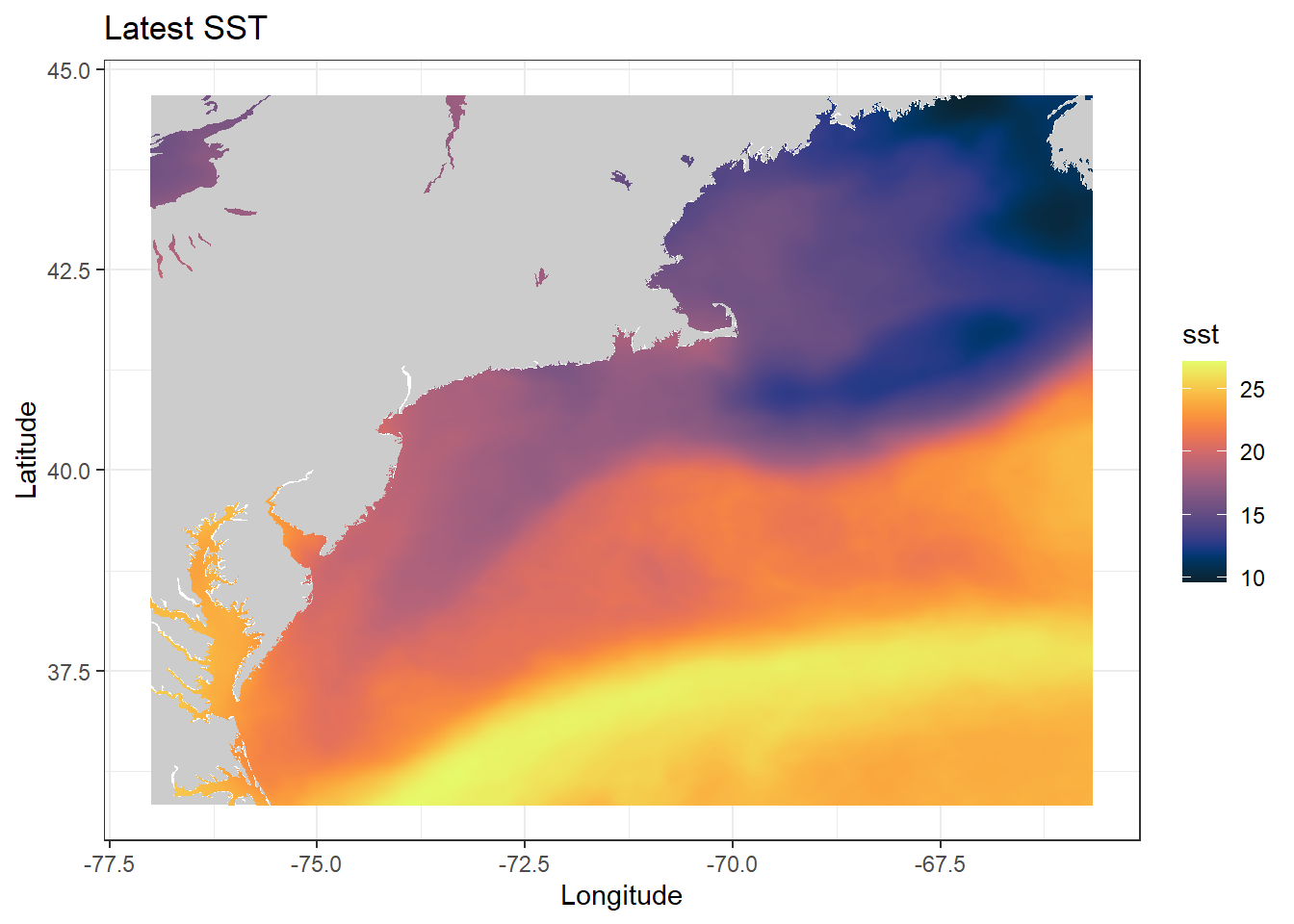
shelf(purrr, raster, sp, tidyr)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.select <- dplyr::select
sst_tbl <- tibble(sst_gd$data) %>%
mutate(
# round b/c of uneven intervals
# unique(sst_gd$data$Lon) %>% sort() %>% diff() %>% table()
lon = round(lon, 2), lat = round(lat, 2), date = as.Date(time, "%Y-%m-%dT00:00:00Z")) %>%
select(-time) %>%
filter(!is.na(sst)) # 13M to 8.8M rows
sst_tbl_mo <- sst_tbl %>%
nest(data = c(lat, lon, sst)) %>%
mutate(raster = purrr::map(data, function(x){
#browser()
sp::coordinates(x) <- ~ lon + lat
sp::gridded(x) <- T
raster::raster(x)
}))
sst_stk <- raster::stack(sst_tbl_mo$raster)
names(sst_stk) <- strftime(sst_tbl_mo$date, "sst_%Y.%m")
raster::crs(sst_stk) <- 4326shelf(stringr)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.epu_sst_avg <- raster::extract(sst_stk, epu_sf, fun = mean, na.rm = T)
epu_sst_sd <- raster::extract(sst_stk, epu_sf, fun = sd, na.rm = T)
epu_sst_tbl <- rbind(epu_sst_avg %>%
as_tibble() %>%
cbind(EPU = epu_sf$EPU, stat = "mean") %>%
pivot_longer(-c(EPU, stat)),
epu_sst_sd %>%
as_tibble() %>%
cbind(EPU = epu_sf$EPU, stat = "sd") %>%
pivot_longer(-c(EPU, stat))) %>%
mutate(EPU = as.character(EPU), date = as.double(str_replace(name, "sst_", ""))) %>%
select(-name) %>%
pivot_wider(names_from = EPU, values_from = value)shelf(dygraphs)##
## The 'cran_repo' argument in shelf() was not set, so it will use
## cran_repo = 'https://cran.r-project.org' by default.
##
## To avoid this message, set the 'cran_repo' argument to a CRAN
## mirror URL (see https://cran.r-project.org/mirrors.html) or set
## 'quiet = TRUE'.epu_sst_tbl %>%
filter(stat == "mean") %>%
select(-stat) %>%
dygraph()